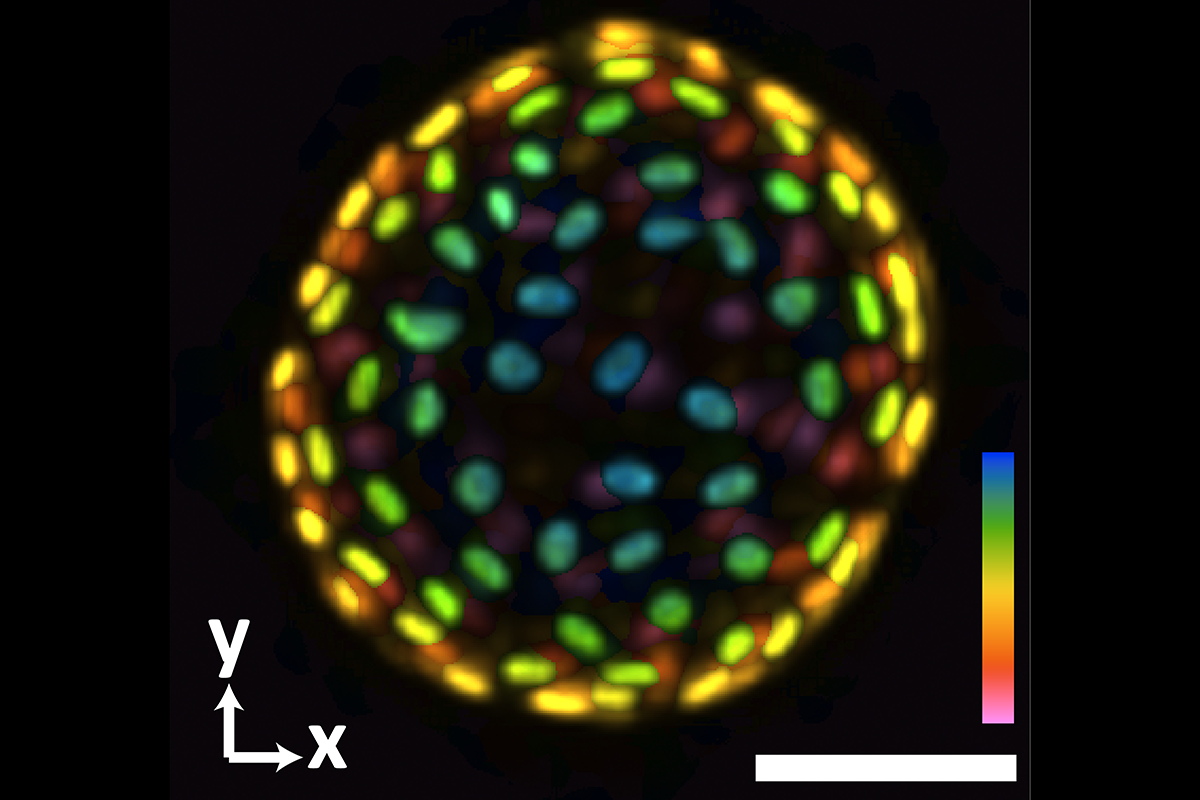
An image of a colon organoid captured by a new system developed in Shu Jia’s lab that can capture dynamic, 3D information about the lab-grown cultures of tissues in a single image. The raw image was captured in 0.1 seconds. The coloration represents depths of 60 and -60 micrometers from the focal plan to depict the organoid in three dimensions. (Image Courtesy: Shu Jia & Wenhao Liu)
Biomedical researchers develop and use organoids as a tool for studying human development and disease. These little lab-grown cultures mimic human organs and provide a sharp view of tissue development, drug interaction, and other biochemical functions, offering an innovative approach to personalized medicine.
“Getting detailed 3D images of these miniature models of organs, and getting a good look at how they change under different conditions or stimulation, can tell us a lot about how the body works,” said Shuichi Takayama, professor and Price Gilbert Jr. Chair in Regenerative Engineering and Medicine in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University. “It can tell us how diseases progress, or how mechanical forces and certain drugs may change or affect cellular behavior.”
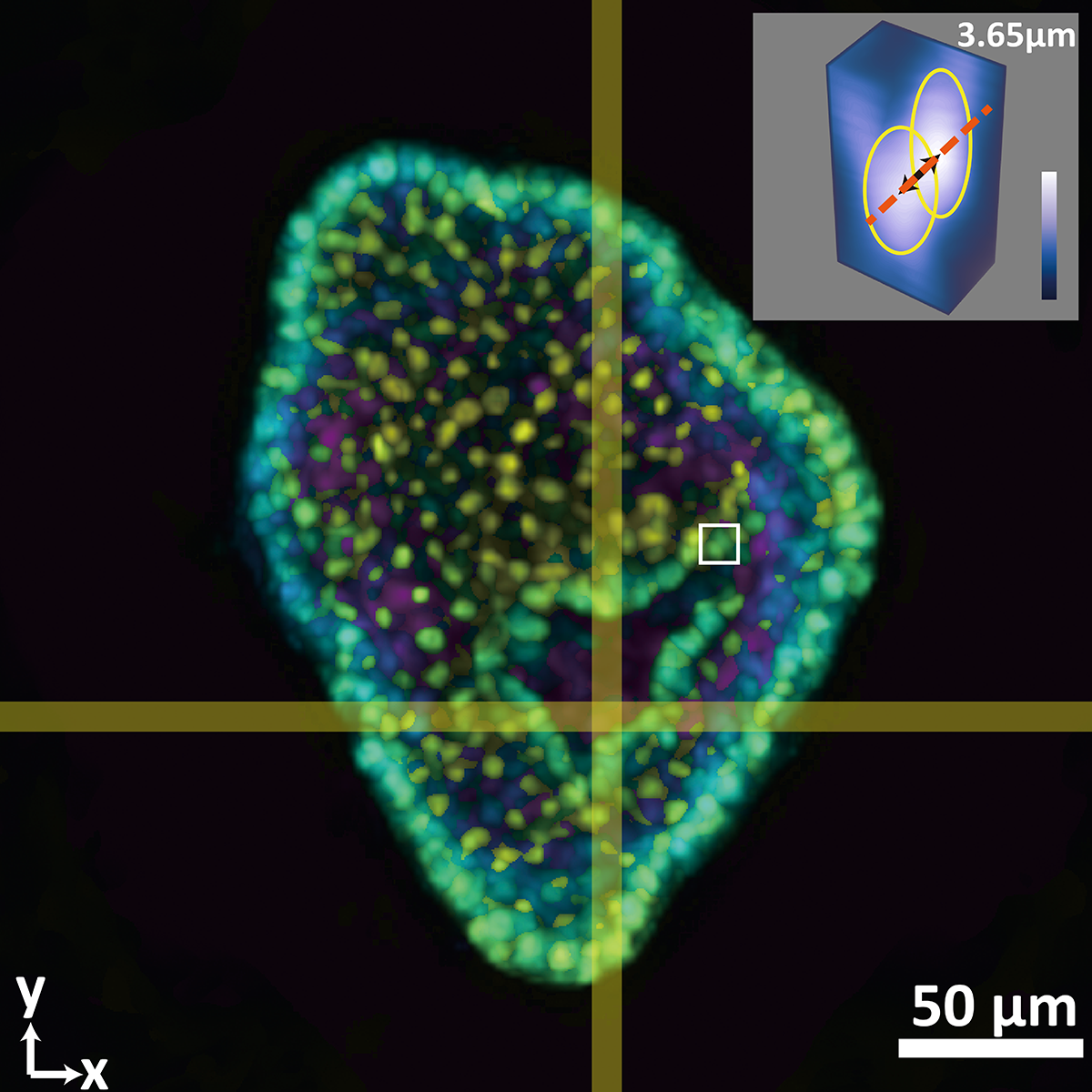
This composite image shows a colon organoid with stained cell nuclei reconstructed from a raw image taken by a new system developed in the Coulter Department. The large image shows depth color coded from yellow (-56.2 micrometers from the focal plane) to purple (+56.2 micrometers from the focal plane). The inset shows a tight crop of the distance between two cells of 3.65 micrometers. The single raw image was captured in 0.1 seconds. (Image Courtesy: Shu Jia & Wenhao Liu)
The trick is in getting those detailed images. Fluorescence 3D microscopy has helped transform the study of organoids at the cellular and subcellular levels — though with a few drawbacks. Conventional methods are time consuming and don’t adequately capture the fast, dynamic, sometimes unpredictable cellular and tissue processes of these model systems.
Now, a team of Georgia Tech researchers has built a better system to quickly produce high-resolution 3D images in real time, providing a quantitative analysis of organoids. Led by Coulter BME Assistant Professor Shu Jia, their custom-built microscope can reconstruct a comprehensive 3D representation with a single camera image. They described their system in the journal Biosensors and Bioelectronics.
Jia’s new system builds on his lab’s growing body of work in next-generation imaging systems. Conventional 3D imaging technologies rely on time-consuming, redundant scanning-based techniques, which can result in damaged cells and compromised images. Jia’s team has pioneered a faster light-field system that provides greater resolution and minimizes photo damage. Their new system does all of that and more.
“This latest system is novel because it is entirely custom-built for imaging at the tissue and animal scale,” said Jia, who earlier this year received a CAREER award from the National Science Foundation. “We built everything from scratch on an optical table.”
Adding a hybrid point-spread function to the new system allows researchers to capture scanning-free recordings of intact organoids in all of their dynamic glory in milliseconds instead of minutes or even hours using conventional methods. With a single camera image, Jia’s system can reconstruct a time-lapse observation of the 3D volume of the samples.
“We can look — cell by cell — throughout the entire organoid, in high spatial and time resolution, and see from multiple angles what happens as a result of an external perturbation, or a response to a specific drug, or any change in the overall environment,” Jia said.
He said the imaging systems coming out of his lab have the potential to transform conventional 3D microscopy.
“Because this is a custom-built system, it’s very flexible and adaptive,” he added. “It works with organoids, but similarly, it can work with animal models. I think we can extend this method to different areas of research. There are a number of potential collaborations we are exploring.”
This research was supported by the National Institutes of Health (grant Nos. R35GM124846 and AI116482) and the National Science Foundation (grant Nos. EFMA1830941 and 2145235). Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of any funding agency.
Latest BME News
Jo honored for his impact on science and mentorship
The department rises to the top in biomedical engineering programs for undergraduate education.
Commercialization program in Coulter BME announces project teams who will receive support to get their research to market.
Courses in the Wallace H. Coulter Department of Biomedical Engineering are being reformatted to incorporate AI and machine learning so students are prepared for a data-driven biotech sector.
Influenced by her mother's journey in engineering, Sriya Surapaneni hopes to inspire other young women in the field.
Coulter BME Professor Earns Tenure, Eyes Future of Innovation in Health and Medicine
The grant will fund the development of cutting-edge technology that could detect colorectal cancer through a simple breath test
The surgical support device landed Coulter BME its 4th consecutive win for the College of Engineering competition.